
Tumor necrosis factor-related lncRNAs predict prognosis and immunotherapy response for patients with lung adenocarcinoma
Highlight box
Key findings
• We constructed and validated a TNF-related lncRNA signature to predict prognosis and immunotherapy response for patients with LUAD in this study.
What is known and what is new?
• The TNF family is involved in tumorigenesis and tumor progression. Various lncRNAs play important roles by mediating TNF family members in cancers. However, the clinical value of TNF-related lncRNAs in LUAD is still not known.
• We constructed and validated a TNF-related lncRNA signature to predict prognosis and immunotherapy response in LUAD.
What is the implication, and what should change now?
• This study indicated that TNF-related lncRNAs showed good performance to predict prognosis and immunotherapy response in LUAD. However, more in vitro and/or in vivo experiments are required to verify the conclusion.
Introduction
Lung cancer is the most common cause of cancer death with an increasing annual incidence rate (1). It mainly includes non-small cell lung cancer (NSCLC) and small cell lung cancer (SCLC), with NSCLC accounting for about 80–85% all cases of lung cancer. Thereinto, more than 50% of NSCLC patients are classified as lung adenocarcinoma (LUAD) (2,3). Although drugs targeting driver genes of LUAD have improved clinical treatment, LUAD patients inevitably develop drug resistance after treatment with these targeted drugs (4). With the development of the immune check-point inhibitors (ICIs) targeting programmed cell death protein 1 (PD-1), programmed death ligand-1 (PD-L1), and cytotoxic T-lymphocyte-associated antigen 4 (CTLA-4), immunotherapy has revolutionized LUAD treatment in the recent decade and provided new approaches to improve the survival of LUAD patients (5,6). However, only a small number of LUAD patients respond to immunotherapy (7). Therefore, there is an urgent need to discover novel biomarkers to predict the treatment response and prognosis for LUAD patients.
The tumor necrosis factor (TNF) family consists of a 19 TNF ligands superfamily (TNFSF) and a 29 TNF receptor superfamily (TNFRSF). The TNF family plays an important role in tumorigenesis and tumor progression (8). Studies have reported that a TNF family-based signature can efficiently predict the prognosis and response to immunotherapy and chemotherapy in several cancers, including LUAD, SCLC, and colorectal cancer (9-11). Moreover, the TNFSF/TNFRSF system has been shown to mediate inflammation-related pathways that are involved in many steps of the immune response (12), highlighting that anti-TNF therapy can effectively relieve inflammation-related autoimmune diseases, such as rheumatoid arthritis and inflammatory bowel disease (8), and cancer pain (13). Therefore, the therapeutic approaches to intervene TNF-related signaling pathways may have a lot of potential.
Long non-coding RNAs (lncRNAs) are RNAs with little or no protein-coding function consisting of more than 200 nucleotides (14). They are involved in drug resistance, carcinogenesis, and inflammatory and immune responses by interacting with RNA-binding proteins and transcription factors (15-17). Studies identified the dual role of lncRNAs by interacting various signal pathways in LUAD. Some lncRNAs promoted the progression of LUAD, such as lncRNA PCAT6 by sponging miR-545-3p and lncRNA FOXD2-AS1 by regulating cell cycle (18,19). However, some lncRNAs inhibited the progression of LUAD, such as lncRNA SNHG5 by regulating epithelial-mesenchymal transition and lncRNA HITT by attenuating Rab5-mediated endocytosis (20,21). Various lncRNAs have been demonstrated to play their roles by mediating TNF family members in multiple cancers, including lung cancer (22-26). Therefore, TNF-related lncRNAs may be a promising research direction in understanding the treatment and prognosis of cancers. However, the expression patterns and clinical significance of TNF-related lncRNAs in LUAD are still not fully clear.
In this study, we systematically explored the expression patterns and clinical significance of TNF-related lncRNAs using RNA-seq data of LUAD patients from The Cancer Genome Atlas (TCGA). We constructed a TNF-related lncRNA prognostic signature by the univariate Cox proportional hazards regression and least absolute shrinkage and selection operator (LASSO)-Cox proportional hazards regression method in the training dataset. Then, the signature was validated in the testing dataset and the entire patient dataset. Finally, we further explored the signature-related biological pathways and the correlation of the signature and immunotherapy response. We present the following article in accordance with the TRIPOD reporting checklist (available at https://jtd.amegroups.com/article/view/10.21037/jtd-23-184/rc).
Methods
Data collection
We collected RNA-seq data of 539 LUAD patients and 59 healthy controls with the corresponding clinical data, including age, gender, and tumor-node-metastasis (TNM) stage, and overall survival (OS) from the TCGA database (http://www.ncbi.gov/repository). Patients who had incomplete clinical data and OS <30 days were excluded, thus 493 LUAD patients were eventually included in the study. The enrolled 493 LUAD patients were randomly assigned into a training dataset and a testing dataset in a 1:1 ratio. Thus, the entire patient dataset, training dataset, and testing dataset consisted of data from 493 patients, 247 patients, and 246 patients, respectively. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013).
Establishment and validation of TNF family-related lncRNA prognostic signature
In training dataset, Pearson correlation analysis was used to identify the correlation of lncRNAs and TNF family genes. The univariate Cox proportional hazards regression analysis was used to identify TNF-related lncRNAs that were related with survival of LUAD patients. With the selected genes, the LASSO-Cox proportional hazards regression analysis was used to construct a TNF-related lncRNA prognostic signature. The risk scores were calculated based on the LASSO regression coefficients. According to the median risk score, these enrolled patients were divided into high- and low-risk subgroups. Then, the Kaplan-Meier (KM) survival analysis was used to evaluate the prognostic value of the signature. To identify the prognostic robustness of the signature from the training dataset, we further assessed its performance in the testing dataset and the entire patient dataset. Finally, univariate and multivariate Cox regression analyses were used to evaluate whether the signature can independently predict prognosis by analyzing the correlation of the signature and traditional clinical factors, including age, gender, and stage.
Functional enrichment analysis
To identify the signature related biological pathways, we used Gene Ontology (GO) functional annotation and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis to explore the potential molecular mechanisms based on differentially expressed genes (DEGs) between the high- and low-risk groups in the clusterProfiler package in R (R Foundation for Statistical Computing, Vienna, Austria).
Tumor immune dysfunction and exclusion analysis
The tumor immune dysfunction and exclusion (TIDE) algorithm was used to predict immune escape and clinical responses to immune checkpoint blockade. Patients with low TIDE prediction scores were deemed to have a good clinical effect to immunotherapy and no immune escape, whereas patients with high TIDE prediction scores tend to be non-responders and have immune escape.
Statistical analysis
R language (version 4.0.0) was the main software tool used for statistical analysis. Univariate, LASSO-Cox, and multivariate Cox regression analyses were used to explore the prognostic value of the signature and the correlation of the signature and traditional clinical factors. KM survival analysis was used to evaluate the survival status.
Time-dependent area under the receiver operating characteristic (ROC) curve (AUC) was used to assess the predictive value of the signature to 1-, 2-, and 3-year OS. Student’s t-test was used to analyze significance among variables. P value <0.05 was considered statistically significant.
Results
Identification of the prognostic value of TNF family-related lncRNAs in LUAD patients
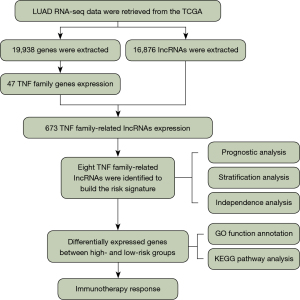
The flowchart of this study is shown in Figure 1. In this study, we enrolled 47 TNF family genes and 16,876 lncRNAs by retrieving the RNA-seq data of LUAD patients from TCGA (Figure 1). Pearson correlation analysis further demonstrated that the expression of 673 lncRNAs was closely related with TNF family genes (|Pearson R|>0.4, P<0.001). Then, the univariate Cox proportional hazards regression analysis was used to evaluate the prognostic value of the identified TNF-related lncRNAs in LUAD. A total of 63 TNF-related lncRNAs significantly associated with OS were identified (P<0.05). After LASSO analysis, 8 TNF-related lncRNAs obviously related with OS were acquired to construct the risk signature (Figure 2). Among these 8 genes, 4 genes (LINC01137, LINC02310, AC017100.1, and AC108676.1) were confirmed as risky factors with hazard ratios (HRs) >1, whereas 4 genes (AC118757.1, AL357873.1, AC104971.3, and HLA-DQB1-AS1) were confirmed as protective factors with HRs <1 (Figure 2).
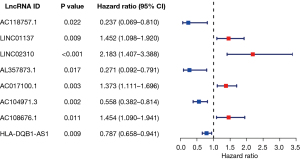
Additionally, KM survival analysis showed that patients with high expression status of 4 genes (LINC01137, LINC02310, AC017100.1, and AC108676.1) had shorter survival time than those with low expression status, whereas patients with high expression of 4 genes (AC118757.1, AL357873.1, AC104971.3, and HLA-DQB1-AS1) had longer survival than those with low expression (P<0.05, Figure 3).
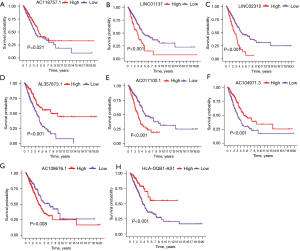
Establishment of a TNF family-related lncRNA prognostic signature using the LUAD patients in the training dataset
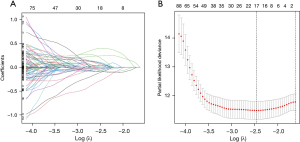
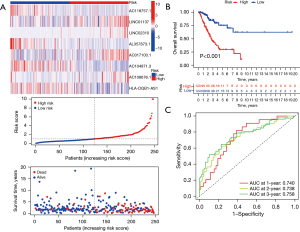
After univariate Cox proportional hazards regression analysis, 8 TNF-related lncRNAs were selected to construct the signature by LASSO-Cox proportional hazards regression analysis in the training dataset (Figure 4). The risk formula was established with the enrolled 8 TNF-related lncRNAs: risk score = (−1.18× AC118757.1) + (0.28× LINC01137) + (0.55× LINC02310) + (−1.21× AL357873.1) + (0.38× AC017100.1) + (−0.45× AC104971.3) + (0.31× AC108676.1) + (−0.15× HLA-DQB1-AS1). The gene expression profiles, the corresponding risk scores, and survival status of the identified 8 TNF-related lncRNAs are displayed in Figure 5A. According to the median risk score, these enrolled patients were divided into high- and low-risk subgroups. Moreover, the KM survival analysis indicated that patients in the high-risk group showed significantly less favorable OS than that of low-risk group (Figure 5B). Subsequently, time-dependent AUC values were calculated to evaluate the ability of the TNF-related lncRNA prognostic signature to predict 1-, 2-, and 3-year OS in the training dataset. These values were 0.740, 0.738, and 0.758, respectively (Figure 5C).
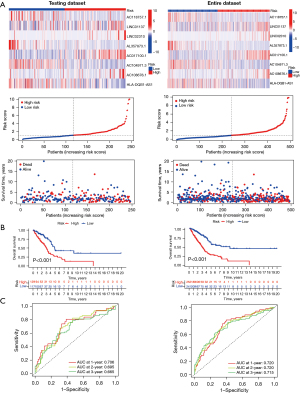
Validation of the TNF family-related lncRNA prognostic signature
To identify the prognostic robustness of the TNF-related lncRNA prognostic signature from the training dataset, we further assessed its performance in the testing dataset and the entire patient dataset. The gene expression profiles, the corresponding risk scores, and survival status in the testing dataset and entire patient dataset are displayed in Figure 6A. As expected, the KM survival analysis showed that patients in the high-risk group showed significantly less favorable OS than that of low-risk group in the testing dataset (P<0.001) and entire patient dataset (P<0.001) (Figure 6B). Time-dependent AUC values were also used to assess the predictive value the signature to 1-, 2-, and 3-year survival. These values were 0.706, 0.695, and 0.665 in the testing dataset, and 0.720, 0.720, and 0.715 in entire patient dataset, respectively (Figure 6C). Therefore, these results indicated that TNF-related lncRNA prognostic signature had a good prognostic predictive value for LUAD patients.
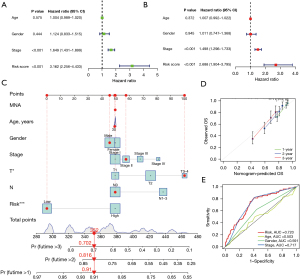
Evaluation of the performance of TNF-related lncRNA prognostic signature in different clinical subgroups
To further verify the prognostic value of TNF family-related lncRNA prognostic signature in different clinical subgroups, the univariate and multivariate Cox regression analysis were performed. The results showed that stage and risk score were independent prognostic factors (Figure 7A,7B). Then, the nomogram was constructed to quantitatively evaluate the survival probability with traditional clinical variables and the risk score (Figure 7C). The calibration diagram showed that the prediction and observation values of nomogram model were consistent (Figure 7D). Moreover, the AUCs showed that the AUC value of risk score (AUC =0.720) was higher than that of age (AUC =0.503) and gender (AUC =0.591) (Figure 7E).
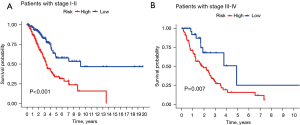
The treatment regimen and prognosis are significantly different between different cancer stages, so we further evaluated the prognostic value of the TNF-related lncRNA prognostic signature in different clinical stages. The KM survival analysis showed that the early-stage (stages I and II) and advanced-stage (stages III and IV) patients had less favorable OS in the high-risk group than that of the low-risk group (Figure 8, P<0.001). The results showed that the signature maintained a reliable predictive performance in different subgroups of LUAD patients.
Identification of TNF-related lncRNA prognostic signature-related biological pathways
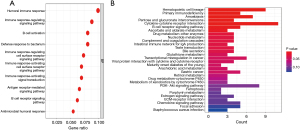
As the TNF-related lncRNA prognostic signature had a high prognostic predictive value for LUAD patients, we continued to explore the biological features. The GO and KEGG pathway analysis were used to explore the signature-related biological pathways. The results of GO analysis indicated that these genes were closely involved in the biological process of humoral immune response, immune response-related signaling pathway, B cell activation, and B cell receptor signaling pathway (Figure 9A). Meanwhile, KEGG analysis showed that primary immunodeficiency, cytokine-cytokine receptor interaction, B cell receptor signaling pathway, complement, and coagulation cascades were mainly enriched (Figure 9B).
Correlation of TNF-related lncRNA prognostic signature and immunotherapy response
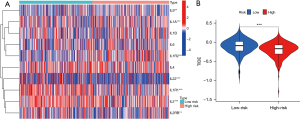
As the results of GO and KEGG pathway analysis showed that it was closely relevant to the immune-related signaling pathway, we further evaluated the correlation of the TNF-related lncRNA prognostic signature and immunotherapy response. The results showed that many interleukin (IL) subtypes, including IL1A, IL1R1, IL1R2, IL2, IL2RB, IL9, and IL22, were closely associated with the risk scores (Figure 10A). Then, we further scored the TIDE, and found that high-risk patients had a lower TIDE score than that of low-risk patients, indicating that high-risk patients identified by TNF-related lncRNA prognostic signature may be appropriate candidates for immunotherapy (Figure 10B).
Discussion
LUAD, as the primary subtype of NSCLC, has become one of the most lethal cancers. With increasing numbers of the identified biomarkers in recent years, we have a better understanding of the origin of LUAD to improve the treatment and prognosis, but the recurrence and the survival rates remain unfavorable (27,28). Therefore, looking for better predictive biomarkers of treatment response and prognosis would be helpful for optimization of personalized treatment and prognosis management of LUAD patients. The TNF family is expressed in almost all cells and is involved in many cellular processes, including cell survival and immunity (29). Studies have demonstrated that the TNF-related signature can predict the prognosis of various cancers, such as cervical cancer, bladder cancer, and SCLC (10,30,31). Moreover, various lncRNAs have been demonstrated to play important roles by mediating TNF family members in multiple cancers, including lung cancer (22-25). These TNF-related lncRNAs were involved in carcinogenesis, and inflammatory and immune responses (22-25). A number of studies have also demonstrated that some of TNF-related lncRNAs are closely associated with cancer proliferation and survival (32-38). Thus, we analyzed the expression patterns and clinical significance of TNF-related lncRNAs in LUAD. In this study, we identified 8 TNF-related lncRNAs that were closely associated with OS of LUAD patients by univariate Cox proportional hazards regression analysis. Then, we used the identified 8 lncRNAs to construct TNF-related lncRNA prognostic signature, and found that the signature had a high predictive capacity of prognosis for LUAD in 3 datasets. Furthermore, the univariate and multivariate Cox regression analysis showed that stage and risk score were independent prognostic factors in different clinical subgroups, and the KM survival analysis also showed that the early-stage (stages I and II) and advanced-stage (stages III and IV) patients had less favorable OS in the high-risk group than that of the low-risk group.
Studies have reported that the enrolled 8 TNF-related lncRNAs (LINC01137, LINC02310, AC017100.1, AC108676.1, AC118757.1, AL357873.1, AC104971.3, and HLA-DQB1-AS1) are closely related with cancer progression. LINC01137 has been shown to play an oncogenic role in NSCLC and oral squamous cell carcinoma, which is similar with our results that NSCLC patients with low expression of LINC01137 were likely to exhibit better outcomes (32,33). Zhao et al. demonstrated that LINC02310, as an enhancer in LUAD, obviously promoted the growth and proliferation of LUAD cells, and its overexpression caused low survival probability (34). In addition, AC017100.1 was shown to be a protective factor for the prognosis of prostate cancer (35). HLA-DQB1-AS1 has been shown to interact with zinc finger RANBP2-type containing 2 (ZRANB2) protein to promote proliferation and inhibit apoptosis in hepatocellular carcinoma (36), suggest that higher expression of HLA-DQB1-AS1 might have a negative impact on survival in hepatocellular carcinoma, while it was demonstrated to be beneficial for the survival of patients with LUAD and melanoma (37,38). Therefore, the role of HLA-DQB1-AS1 in cancer was related to the type of cancers, and the mechanism needs to explore further. Moreover, AC108676.1, AC118757.1, AL357873.1, and AC104971.3 have not been previously reported. This is the first time that they have been reported to be considered as the prognostic biomarkers of LUAD.
The GO and KEGG pathway analysis provided a further understanding of TNF family-related lncRNA prognostic signature-related biological pathways, and found that these genes were closely involved in immune-related signaling pathways. Immunotherapy has emerged as a novel treatment modality and become the standard treatment for many kinds of cancers, including LUAD (39). However, some LUAD patients still develop disease progression (40). Many factors have been revealed to affect the response to ICIs, such as the pro-inflammatory cytokines and the expression level of PD-L1 and others (41,42). In this study, we found that many IL subtypes, including IL1A, IL1R1, IL1R2, IL2, IL2RB, IL9, and IL22, were closely associated with the risk scores, suggesting that these pro-inflammatory cytokines are involved in immunotherapy response of LUAD patients. Moreover, studies have revealed 2 distinct mechanisms of tumor immune evasion: T cell dysfunction and T cell exclusion (43,44). From this, Jiang et al. developed TIDE to identify factors that underlie these 2 mechanisms (45). Compared with widely used ICI response biomarkers, such as PD-L1 level and tumor mutation load, TIDE signature showed better predictor for both anti-PD1 and anti-CTLA4 therapies (45). Thus, we further scored the TIDE, and found that high-risk patients had a lower TIDE score than that of low-risk patients, indicating that high-risk patients identified by TNF-related lncRNA prognostic signature may be appropriate candidates for immunotherapy.
However, this study has some limitations. We only used the data from the public dataset TCGA to construct and validate TNF-related lncRNA prognostic signature. More in vitro and/or in vivo experiments are needed to verify the clinical predictive ability of the signature.
Conclusions
In conclusion, this study is the first to construct and validate a prognostic predictive signature of LUAD patients based on TNF-related lncRNAs. The signature showed good performance to predict immunotherapy response in LUAD. Therefore, this signature may provide new strategies for individualized treatment of LUAD patients.
Acknowledgments
Funding: This study was supported by the Tackling-plan Project of Henan Department of Science and Technology (No. 212102310325) and the Tackling-plan Project of Henan Medical Science and Technology (No. LHGJ20200164).
Footnote
Reporting Checklist: The authors have completed the TRIPOD reporting checklist. Available at https://jtd.amegroups.com/article/view/10.21037/jtd-23-184/rc
Peer Review File: Available at https://jtd.amegroups.com/article/view/10.21037/jtd-23-184/prf
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://jtd.amegroups.com/article/view/10.21037/jtd-23-184/coif). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Sung H, Ferlay J, Siegel RL, et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2021;71:209-49. [Crossref] [PubMed]
- Travis WD, Brambilla E, Nicholson AG, et al. The 2015 World Health Organization classification of lung tumors: impact of genetic, clinical and radiologic advances since the 2004 classification. J Thorac Oncol 2015;10:1243-60. [Crossref] [PubMed]
- Davidson MR, Gazdar AF, Clarke BE. The pivotal role of pathology in the management of lung cancer. J Thorac Dis 2013;5:S463-78. [PubMed]
- Barbar J, Armach M, Hodroj MH, et al. Emerging genetic biomarkers in lung adenocarcinoma. SAGE Open Med 2022;10:20503121221132352. [Crossref] [PubMed]
- Zhan X, Feng S, Zhou X, et al. Immunotherapy response and microenvironment provide biomarkers of immunotherapy options for patients with lung adenocarcinoma. Front Genet 2022;13:1047435. [Crossref] [PubMed]
- Huang Z, Zhou C, Xiong Y, et al. PD-1 inhibitor versus bevacizumab in combination with platinum-based chemotherapy for first-line treatment of advanced lung adenocarcinoma: A retrospective-real world study. Front Oncol 2022;12:909721. [Crossref] [PubMed]
- Passaro A, Brahmer J, Antonia S, et al. Managing resistance to immune checkpoint inhibitors in lung cancer: treatment and novel strategies. J Clin Oncol 2022;40:598-610. [Crossref] [PubMed]
- Croft M, Benedict CA, Ware CF. Clinical targeting of the TNF and TNFR superfamilies. Nat Rev Drug Discov 2013;12:147-68. [Crossref] [PubMed]
- Zhang C, Zhang G, Sun N, et al. Comprehensive molecular analyses of a TNF family-based signature with regard to prognosis, immune features, and biomarkers for immunotherapy in lung adenocarcinoma. EBioMedicine 2020;59:102959. [Crossref] [PubMed]
- Zhang Z, Wu P, Zhang C, et al. Tumor necrosis factor family member profile predicts prognosis and adjuvant chemotherapy benefit for patients with small-cell lung cancer. Front Immunol 2021;12:745769. [Crossref] [PubMed]
- Xiao Z, Nie K, Han T, et al. Development and validation of a TNF family-based signature for predicting prognosis, tumor immune characteristics, and immunotherapy response in colorectal cancer patients. J Immunol Res 2021;2021:6439975. [Crossref] [PubMed]
- Wallach D. The tumor necrosis factor family: family conventions and private idiosyncrasies. Cold Spring Harb Perspect Biol 2018;10:a028431. [Crossref] [PubMed]
- Liu Y, Gao Y, Lin T. Expression of interleukin-1 (IL-1), IL-6, and tumor necrosis factor-α (TNF-α) in non-small cell lung cancer and its relationship with the occurrence and prognosis of cancer pain. Ann Palliat Med 2021;10:12759-66. [Crossref] [PubMed]
- Nagano T, Fraser P. No-nonsense functions for long noncoding RNAs. Cell 2011;145:178-81. [Crossref] [PubMed]
- Jarroux J, Morillon A, Pinskaya M. History, discovery, and classification of lncRNAs. Adv Exp Med Biol 2017;1008:1-46. [Crossref] [PubMed]
- Zhou H, Feng B, Abudoureyimu M, et al. The functional role of long non-coding RNAs and their underlying mechanisms in drug resistance of non-small cell lung cancer. Life Sci 2020;261:118362. [Crossref] [PubMed]
- Fang Y, Fullwood MJ. Roles, functions, and mechanisms of long non-coding RNAs in cancer. Genomics Proteomics Bioinformatics 2016;14:42-54. [Crossref] [PubMed]
- Yang C, Huang H, Li Y, et al. LncRNA PCAT6 promotes proliferation, migration, invasion, and epithelial-mesenchymal transition of lung adenocarcinoma cell by targeting miR-545-3p. Mol Biol Rep. 2023;50:3557-68. [Crossref] [PubMed]
- Yuan Y, Yu P, Shen H, et al. LncRNA FOXD2-AS1 increased proliferation and invasion of lung adenocarcinoma via cell-cycle regulation. Pharmgenomics Pers Med 2023;16:99-109. [Crossref] [PubMed]
- Li Z, Wu Y, Zhang C, et al. LncRNA SNHG5 suppresses cell migration and Invasion of human lung adenocarcinoma via regulation of epithelial-mesenchymal transition. J Oncol 2023;2023:3335959. [Crossref] [PubMed]
- Wang X, Zheng S, Yang F, et al. lncRNA HITT inhibits metastasis by attenuating Rab5-mediated endocytosis in lung adenocarcinoma. Mol Ther 2022;30:1071-88. [Crossref] [PubMed]
- Xu K, Meng Z, Xian XM, et al. LncRNA PVT1 induces chondrocyte apoptosis through upregulation of TNF-α in synoviocytes by sponging miR-211-3p. Mol Cell Probes 2020;52:101560. [Crossref] [PubMed]
- Tang ZL, Zhang K, Lv SC, et al. LncRNA MEG3 suppresses PI3K/AKT/mTOR signalling pathway to enhance autophagy and inhibit inflammation in TNF-α-treated keratinocytes and psoriatic mice. Cytokine 2021;148:155657. [Crossref] [PubMed]
- Wang Z, Kun Y, Lei Z, et al. LncRNA MIAT downregulates IL-1β, TNF-ɑ to suppress macrophage inflammation but is suppressed by ATP-induced NLRP3 inflammasome activation. Cell Cycle 2021;20:194-203. [Crossref] [PubMed]
- Wang J, Yuan Y, Tang L, et al. Long non-coding RNA-TMPO-AS1 as ceRNA binding to let-7c-5p upregulates STRIP2 expression and predicts poor prognosis in lung adenocarcinoma. Front Oncol 2022;12:921200. [Crossref] [PubMed]
- Tang H, Chen H, Yuan H, et al. Comprehensive analysis of necroptosis-related long noncoding RNA to predict prognosis, immune status, and immunotherapeutic response in clear cell renal cell carcinoma. Transl Cancer Res 2022;11:4254-71. [Crossref] [PubMed]
- Fan XX, Wu Q. Decoding lung cancer at single-cell level. Front Immunol 2022;13:883758. [Crossref] [PubMed]
- Ma B, Geng Y, Meng F, et al. Identification of a sixteen-gene prognostic biomarker for lung adenocarcinoma using a machine learning method. J Cancer 2020;11:1288-98. [Crossref] [PubMed]
- Balkwill F. Tumour necrosis factor and cancer. Nat Rev Cancer 2009;9:361-71. [Crossref] [PubMed]
- Ma Y, Zhang X, Yang J, et al. Comprehensive molecular analyses of a TNF family-based gene signature as a potentially novel prognostic biomarker for cervical cancer. Front Oncol 2022;12:854615. [Crossref] [PubMed]
- Li H, Liu S, Li C, et al. TNF family-based signature predicts prognosis, tumor microenvironment, and molecular subtypes in bladder carcinoma. Front Cell Dev Biol 2021;9:800967. [Crossref] [PubMed]
- Yao Y, Yang F, Chen A, et al. Costimulatory molecule-related lncRNA model as a potential prognostic biomarker in non-small cell lung cancer. Cancer Med 2023;12:6419-36. [Crossref] [PubMed]
- Du Y, Yang H, Li Y, et al. Long non-coding RNA LINC01137 contributes to oral squamous cell carcinoma development and is negatively regulated by miR-22-3p. Cell Oncol (Dordr) 2021;44:595-609. [Crossref] [PubMed]
- Zhao W, Wang J, Luo Q, et al. Identification of LINC02310 as an enhancer in lung adenocarcinoma and investigation of its regulatory network via comprehensive analyses. BMC Med Genomics 2020;13:185. [Crossref] [PubMed]
- Wang K, Zhong W, Long Z, et al. 5-Methylcytosine RNA methyltransferases-related long non-coding RNA to develop and validate biochemical recurrence signature in prostate cancer. Front Mol Biosci 2021;8:775304. [Crossref] [PubMed]
- Long J, Liu L, Zhou X, et al. HLA-DQB1-AS1 promotes cell proliferation, inhibits apoptosis, and binds with ZRANB2 protein in hepatocellular carcinoma. J Oncol 2022;2022:7130634. [Crossref] [PubMed]
- Liu B, Yang S. A five autophagy-related long non-coding RNA prognostic model for patients with lung adenocarcinoma. Int J Gen Med 2021;14:7145-58. [Crossref] [PubMed]
- Li Z, Wei J, Zheng H, et al. The new horizon of biomarker in melanoma patients: A study based on autophagy-related long non-coding RNA. Medicine (Baltimore) 2022;101:e28553. [Crossref] [PubMed]
- Sharma P, Allison JP. The future of immune checkpoint therapy. Science 2015;348:56-61. [Crossref] [PubMed]
- Skoulidis F, Goldberg ME, Greenawalt DM, et al. STK11/LKB1 mutations and PD-1 inhibitor resistance in KRAS-mutant lung adenocarcinoma. Cancer Discov 2018;8:822-35. [Crossref] [PubMed]
- Wen Y, Zhu Y, Zhang C, et al. Chronic inflammation, cancer development and immunotherapy. Front Pharmacol 2022;13:1040163. [Crossref] [PubMed]
- Borghaei H, Paz-Ares L, Horn L, et al. Nivolumab versus docetaxel in advanced nonsquamous non-small-cell lung cancer. N Engl J Med 2015;373:1627-39. [Crossref] [PubMed]
- Gajewski TF, Schreiber H, Fu YX. Innate and adaptive immune cells in the tumor microenvironment. Nat Immunol 2013;14:1014-22. [Crossref] [PubMed]
- Joyce JA, Fearon DT. T cell exclusion, immune privilege, and the tumor microenvironment. Science 2015;348:74-80. [Crossref] [PubMed]
- Jiang P, Gu S, Pan D, et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat Med 2018;24:1550-8. [Crossref] [PubMed]
(English Language Editor: J. Jones)


