
Association of interferon regulator factor 1 upregulation with pulmonary arterial hypertension
Highlight box
Key findings
• Upregulation of interferon regulator factor 1 (IRF1) in lung tissue and pulmonary arteries may influence the onset and progression of pulmonary arterial hypertension (PAH).
What is known and what is new?
• A considerable amount of research has analyzed the differences in the messenger RNA expression between patients with PAH and healthy controls.
• Our study analyzed Gene Expression Omnibus datasets and found IRF1 to be associated with PAH. We further conducted animal experiments to verify this finding.
What is the implication, and what should change now?
• IRF1 could serve as a therapeutic target for PAH, but further experiments are needed to establish a causal relationship between IRF1 and PAH.
Introduction
Pulmonary arterial hypertension (PAH) refers to a severe and complex disease state that is associated with a poor prognosis characterized by increased pulmonary arterial pressure and elevated pulmonary vascular resistance (PVR) due vasoconstriction and vascular remodeling (1). Pulmonary arterial smooth muscle cells (PASMCs) and pulmonary arterial endothelial cells (PAECs) play crucial roles in the increase in PVR (2). However, the precise nature of the mechanisms contributing to the elevation in PVR remains largely unknown. Data from the National Center for Health Statistics indicate that the 1-year survival rate for patients with untreated PAH is approximately 68% and falls to below 35% at 5 years (3). Even with dedicated clinical treatment, the 1-year survival rate still ranges from 50% to 86% worldwide (4). Additionally, delays in the diagnosis and treatment of PAH are common, as according to the latest European Society of Cardiology and the European Respiratory Society guidelines, PAH should be diagnosed through right-heart catheterization (5,6). Currently, PAH therapy mainly consists of oral administration of endothelin receptor antagonists and phosphodiesterase 5 inhibitors (7). However, more comprehensive investigations are required to identify other avenues of clinical diagnosis and intervention.
Analysis of differentially expressed genes (DEGs) in lung tissue and cells may offer insights into the genes involved in vasoconstriction and vascular remodeling in patients with PAH (8). Stearman et al. (9) used single-cell RNA-sequencing technology to identify genes that were differentially expressed in patients with PAH. Meanwhile, Zheng et al. (10) used several network analysis algorithms to identify key upregulated genes. He et al. (11) analyzed several datasets, reporting a difference in the messenger RNA (mRNA) expression of lung tissue between PAH samples and those of healthy controls. However, experimental validation is needed to confirm the reliability of these analytical results (12).
In this study, we aimed to identify several hub genes by reanalyzing RNA-sequencing data from patients with PAH and healthy controls. We ultimately verified expression changes of interferon regulator factor 1 (IRF1) in animal models. We present this article in accordance with the ARRIVE reporting checklist (available at https://jtd.amegroups.com/article/view/10.21037/jtd-2025-390/rc).
Methods
Data collection
The GSE144274 and GSE243193 datasets from the Gene Expression Omnibus (GEO) database (http://www.ncbi.nlm.nih.gov/geo/) were examined. GSE144274 includes 4 PASMC samples from patients with PAH and from 4 healthy individuals. The platform for GSE144274 is GPL20301 (13). GSE243193 is a GPL16791 Illumina HiSeq 2500-derived human gene data set that includes 55 PAEC samples from both patients with PAH and healthy controls (14). A workflow summarizing the analysis of this study is provided in Figure 1.
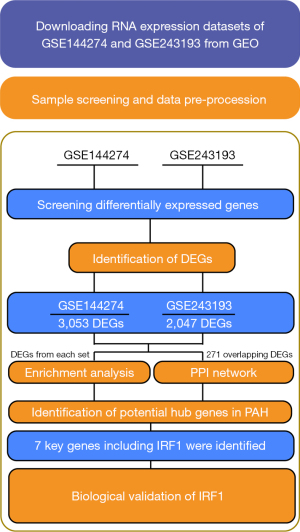
DEG analysis
With the aim of data standardization, we applied log2-transformation and quantile normalization, along with background correction, to raw expression materials using the “limma” package (version 3.52.2) in R software (version 4.2.3; The R Foundation for Statistical Computing). A cutoff value of an adjusted P value <0.05 and |log2 fold change (FC)| >1 was applied to identify DEGs according to log2FC expression. The heatmaps of DEGs and volcano plots were drawn using “ggplot2” packages (version 4.1.2). The overlapping DEGs of GSE144274 and GSE243193 were identified and visualized as Venn diagrams.
Function enrichment analysis
R software (version 4.4.1) was used to perform Gene Ontology (GO) analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis, including for molecular functions (MFs), cellular components (CCs), and biological processes (BPs). A P value less than 0.05 was considered to indicate significant enrichment.
Protein-protein interaction (PPI) network construction
A PPI network was established using Search Tool for the Retrieval of Interacting Genes/Proteins (STRING) online analysis software (https://cn.string-db.org/). Based on the STRING analysis, the MCODE plug-in Cytoscape (version 3.7.1) and three algorithms [maximum neighborhood component (MNC), gene connection degree, and maximal clique centrality (MCC)] were used to screen for the main modules and to identify the hub genes.
A Sugen 5416- and hypoxia-induced PAH mouse model
C57BL/6J male mice (8 to 10 weeks old, weighing 20–25 mg) were obtained from the Soochow University Laboratory Animal Center (SULAC). Using simple randomization method, we divided the mice into a control group (n=12), hypoxia plus Sugen 5416 group (HySu; n=12), and HySu plus 10-hydroxydecanoic acid (10-HDAA; Sanying, Wuhan, Hubei, China) group (HySu + 10-HDAAl n=6). Mice from both the PAH and PAH + 10-HDAA groups were raised in a hypoxic environment (10% O2) and received a subcutaneous injection of Sugen 5416 (20 mg/kg; MedChem Express, Monmouth Junction, NJ, USA) per week for 4 weeks. The control group received injections of 5% saline with the same volume as that of Sugen 5416 applied each week in the PAH group. Mice from the PAH + 10-HDAA group received a 5-week intragastric administration with 100 mg/kg of 10-HDAA once a day at 1 week before Sugen5 416 injection.
Monocrotaline-induced PAH rat model
Sprague-Daley male rats (8 weeks old, weighing 200–250 g) were acquired from the SULAC. The rats were randomly divided into a monocrotaline (MCT; Sanying, Wuhan, Hubei, China) group (MCT; n=6) and a control group (n=6). The rats and the mice mentioned above were raised in a clean environment, provided sufficient water and food, and housed in conditions of 12 hours of alternating light and dark. Rats from the MCT group were intraperitoneally injected with 60 mg/kg of MCT on the first day. Rats from the control group received the same volume of 5% saline injection. After 28 days, the rats were anesthetized and prepared for right heart catheterization (RHC).
RHC
Mice and rats were anesthetized and then received RHC to measure the right ventricular systolic blood pressure (RVSP). After the measurement, the mice and rats were killed, and their hearts and lung tissues were collected. The ratio of the weight of the right ventricle (RV) to the sum of the weight of the left ventricle (LV) and septum (S) was then calculated [RV/(LV+S)].
RNA extraction and real-time quantitative polymerase chain reaction
Total RNA samples of lung tissues and PASMCs were extracted using TRIzol (Takara Bio, Kusatsu, Japan). The sample concentration was then determined with a NanoDrop 1000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). A PrimeScripts RT Master kit (Takara Bio) was employed for reverse transcription. Quantitative reverse transcription polymerase chain reaction (qRT-PCR) was performed on a Bio-Rad CFX 96 PCR system (Bio-Rad Laboratories, Hercules, CA, USA) with FastStart Universal SYBR Green Master (Rox) (Roche, Basel, Switzerland). The GAPDH gene was used to normalize the expression of other gene targets. The primer pairs were as follows: IRF1 forward, 5'-GGC CGA TAC AAA GCA GGA GAA-3'; IRF1 reverse, 5'-GGA GTT CAT GGC ACA ACG GA-3'; GAPDH forward, 5'-GGT GTG AAC CAT GAG AAG TAT GA-3'; and GAPDH reverse, 5'-GAG TCC TTC CAC GAT ACC AAAG-3'.
Western blot analysis
Radioimmunoprecipitation assay lysis was used to dissolve the lung and artery tissue via protease and phosphatase inhibitors (Beyotime, Haimen, China). After the centrifugation of lysate at 12,000 rpm for 15 minutes, the supernatant was collected. BCA Protein Assay Kit (Thermo Fisher Scientific) was used to measure and determine protein concentrations. Protein samples were boiled in a metal bath for 5 minutes with 5× loading buffer and stored at –20 ℃ for further use. Polyvinylidene difluoride membranes (Beyotime) were blocked with NcmBlot blocking buffer (NCM Biotech, Suzhou, China). The primary antibodies adopted were IRF1 (Proteintech, Rosemont, IL, USA) and GAPDH (Proteintech). Following the incubation and rinsing of primary antibodies, the membranes were treated with a 1-hour incubation at room temperature with horseradish peroxidase-conjugated goat anti-rabbit immunoglobulin G (IgG) (Bioss Inc., Woburn, MA, USA). Strip exposure with a chemiluminescence system (Bio-Rad Laboratories) was analyzed with ImageJ software (National Institutes of Health, Bethesda, MD, USA).
Histopathology
After the measurement of RVSP and RV/(LV+S), the lung tissues were flushed with cold 5% saline until they turned white in color. Tissues were then carefully isolated, immersed into paraformaldehyde, and then embedded in paraffin, after which histological analysis was conducted. Hematoxylin and eosin staining and immunofluorescence were used to evaluate the changes in pulmonary artery morphology. The following primary antibodies were used: anti-IRF1 antibody (Proteintech) and anti-β-actin antibody (Proteintech).
Ethical statement
The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). Experiments were performed under a project license (Lunshen No. 230148) granted by the Animal Ethics Committee of The Fourth Affiliated Hospital of Soochow University and in compliance with Soochow University guidelines for the care and use of animals. A protocol was prepared before the study without registration.
Statistical analysis
Statistical analysis was performed using SPSS 26.0 software (IBM Corp., Armonk, NY, USA). Data are expressed as the mean ± standard deviation (SD). If the data were normally distributed with equal variances between two groups, the t-test was used. For three groups, one-way analysis of variance was used along with pairwise comparisons via the least significant difference t-test. If the data were not normally distributed or had unequal variances, the Wilcoxon rank-sum test or the Kruskal-Wallis test was used. A P value of <0.05 was considered statistically significant.
Results
DEG screening in PAH
The gene expression data sets, GSE144274 and GSE243193, were examined: 3,053 DEGs were identified in GSE144274 and 2,054 in GSE243193. An adjusted P value <0.05 and |log2FC| >1 was deemed suitable for DEG screening. There were 1,444 upregulated DEGs in GSE144274 and 1,394 in GSE243193. Meanwhile, there were 1,609 downregulated DEGs in GSE144274. The volcano plots of DEGs from each data set and Venn diagrams of the DEGs are shown in Figure 2.

Functional analysis of the DEGs
GO and KEGG were applied for the DEGs in each of the data sets. The GO analysis results from DEGs in GSE144274 and DEGs in GSE243193 are presented in terms of BP, MF, and CC in Figure 3A,3B. KEGG pathway analysis revealed that the DEGs in GSE144274 were primarily enriched in biological pathways such as the PI3K/Akt signaling pathway, cell cycle, and cytoskeleton in muscles; meanwhile, DEGs in GSE243193 were primarily enriched in MAPK signaling pathway, Ras signaling pathway, and cell adhesion molecules (Figure 3C,3D).

Identification of the key genes in PAH
A PPI network of the overlapping DEGs extracted from GSE144274 and GSE243193 was established to identify the key genes (Figure 4A). A total of 164 nodes and 305 edges were found. Subsequently, the MCODE plug-in was used within Cytoscape. The top 10 key genes identified by the three algorithms (degree, MNC, and MCC) in CytoHubba and the principal module from MCODE were intersected (Figure 4B-4E). A Venn diagram from the three algorithms is presented in Figure 4F. Seven key hub genes were identified: ACTA2, HLA-DRA, HLA-A, PECAM1, HLA-C, IRF1, and CD74.

Validation of IRF1 expression in PAH animal models
Ultimately, IRF1 was chosen for validation in animal experiments. The HySu mouse model and MCT rat model were established. Compared with the control group, the HySu mouse group demonstrated significantly elevated RVSP (25.83±6.62 vs. 54.57±11.98 mmHg; P<0.001; n=6). The RV/(LV+S) was higher in mouse HySu group than in the control group (39.71%±7.02% vs. 52.53%±9.57%; P=0.03; n=6). The distal pulmonary artery walls were thickened, and the lumina were narrowed in both HySu mouse group and MCT rat group. The RVSP of was significantly higher in the MCT rat group than in the control group (19.87±4.79 vs. 39.72±10.45 mmHg; P=0.002; n=6). Compared with that in the control group, RV/(LV+S) in the MCT rat group was significantly higher (28.78%±4.22% vs. 45.49%±10.48%; P=0.005; n=6). These data indicated that the PAH models were established successfully (Figure 5A-5L). Immunofluorescence staining was used to morphologically observe the expression of IRF1 in rats and mice (Figure 5M). The results of IRF1 expression level are summarized in Figure 6. The protein expression of IRF1 was elevated in the lung tissue of both mice and rats (mice: P<0.001, n=6; rats: P=0.04, n=6) as was the expression level in the pulmonary arteries (mice: P<0.001, n=6; rats: P=0.01, n=6). The expression level of IRF1 from the qRT-PCR results were consistent with the findings for protein expression in pulmonary tissue (mice: P=0.001, n=6: rats: P<0.001, n=6) and arteries (mice: P<0.001, n=6; rats: P=0.003, n=6).


IRF1 inhibition could ameliorate PAH in Sugen 5416- and hypoxia-induced PAH mice
To further clarify the relationship between IRF1 and PAH, we used 10-HDAA to establish a PAH mouse model with the inhibition of IRF1 (Figure 7). The protein expression of IRF1 was higher in the PAH group as compared with the control group (P<0.001; n=6). Compared with that in the PAH group, the protein expression of IRF1 was lower in the PAH + 10-HDAA group (P<0.001; n=6). Compared with that in the control group, the protein expression of IRF1 was higher in the 10-HDAA + HySu group (P<0.001; n=6). The RVSP of the HySu group was elevated compared with that in the control group (28.21±8.41 vs. 57.67±11.02 mmHg; P<0.001; n=6). Among mice, RVSP was lower in the PAH+10-HDAA group than in the PAH group (57.67±11.02 vs. 45.83±4.88 mmHg; P=0.04; n=6). The RVSP of the 10-HDAA + HySu group was elevated compared with that in the control group (45.83±4.88 vs. 28.21±8.41 mmHg; P<0.001; n=6). Finally, the RV/(LV+S) in the HySu group was significantly higher than that of the control group (30.71%±5.83% vs. 56.53%±7.29%; P<0.001; n=6); moreover, it was lower in the 10-HDAA + HySu group than in the HySu group (56.53%±7.29% vs. 48.31%±5.26%; P=0.045; n=6). The RV/(LV+S) in the 10-HDAA + HySu group was significantly higher than that of the control group (48.31%±5.26% vs. 30.71%±5.83%; P<0.001; n=6).

Discussion
In this study, we chose GSE144274 and GSE243193 due to their relevance to our research objectives, the quality of the data, and their availability for public access. These datasets contain pertinent information that aligns with our study’s focus on pulmonary arterial hypertension. While there is few study on GSE243193, GSE144274 has been widely cited in the literature, indicating its credibility and significance in the field. We screened out overlapping DEGs from GSE144274 and GSE243193 and conducted functional analysis to investigate the underlying mechanisms. A PPI network was established to screen out the hub genes and pinpoint the key genes base on the results. The principal findings are as follows: (I) seven key genes (ACTA2, HLA-DRA, HLA-A, PECAM1, HLA-C, IRF1, CD74) were identified between patients with PAH and healthy individuals; (II) the upregulation of IRF1 may play a role in the pathophysiological processes associated with PAH.
IRF1 is an interferon regulator of cell proliferation and immune response, functioning as an activator of genes in relation with both adaptive and innate immune responses (15). It is involved in the transcriptional activation process that responds to bacterial and viral pathogen invasion, playing a role in programmed cell death and DNA damage response (16). IRF1 is expressed in a variety of cell types (17), and IRF1 mRNA level appears to significantly increase in response to inflammation reaction or interferon stimulation (18). Complement DNA (cDNA) transfection experiments have demonstrated that IRF1 can activate the promoters of IFN-α/β (19), and IRF1 cDNA expression may induce endogenous IFN-α and IFN-β gene expression in various cell lines (20). It has also been observed that the upregulation of IRF1 expression can induce inflammatory cell death of PAECs in the airway (21). Another study suggests that IRF1 may play an inhibitory role in the regeneration of the endothelium (22). Research has found that the activation of IRF1 can promote the abnormal differentiation and proliferation of smooth muscle cells by regulating inflammatory responses and apoptosis (23). Furthermore, a recent study has shown that inhibiting IRF1 expression can reduce the abnormal proliferation of smooth muscle cells (24). 10-HDAA has been reported to exert an inhibitory effect on the expression of IRF1 (25). Based on its critical role in immune response and gene regulation, IRF1 was selected as our target (19). A previous study has highlighted its significance in programmed cell death of endothelial cells and smooth muscle cells, which in part aligns with the objectives of our research (21). Considering these findings, we applied 10-HDAA as an inhibitor of IRF1 for the intervention experiments in mice. While our current study did not include this intervention on rats, we recognize the potential benefits of such an experiment in further elucidating the effects of IRF1 in vivo. However, rats usually weigh ten times that of mice. The dosage of 10-HDAA required by gavage in rats is too much to afford, making it difficult to proceed with the experiment.
PAECs and PASMCs play key roles in the pulmonary vascular remodeling (26). In one study that used PASMCs from patients with PAH, the conditioned medium from the coculture of M2 macrophages and PASMCs exhibited a more pronounced effect on exacerbating PASMC migration and proliferation, and thus worsened the PAH condition (27). Wang et al. (28) found that dysfunction of bone morphogenetic protein receptor 2 signaling pathway in PASMCs could impair the contractility of pulmonary vessels and increase proliferation, potentially resulting in persistent pulmonary hypertension. In MCT-induced PAH models, enhanced apoptosis and ferroptosis of PAECs were observed (29). Interestingly, upregulation of IRF1 may aggravate the pyroptosis of PAECs in endothelial cells of vessels (30). Recent single-cell mapping research indicates that targeting inflammatory and immune responses may alleviate severe vascular pathology in PAH (31). These findings suggest that IRF1 may be a key immune-related gene in the progression of PAH.
There are several limitations to our study. First, we did not examine the signaling pathways involving IRF1 in depth, necessitating further experiments to support and validate our results. Second, GSE144274 consists of patients with an idiopathic PAH profile and a relatively clear etiology, while GSE243193 includes patients with different types of PAH. Third, a combined analysis of the two gene expression profiles of PASMCs and PAECs that could facilitate the screening for immune-related genes and genes involved in intercellular interactions was not conducted. Finally, our findings do not sufficiently clarify the in vitro role of IRF1 in PAH, although we did manage to construct both a HySu model and an MCT model. Therefore, more in-depth fundamental experiments are required to characterize the relationship between IRF1 and PAH.
Conclusions
Our findings suggest that the expression level of IRF1 in lung tissue and pulmonary arteries may influence the onset and progression of PAH, indicating that IRF1 could serve as a therapeutic target for PAH.
Acknowledgments
None.
Footnote
Reporting Checklist: The authors have completed the ARRIVE reporting checklist. Available at https://jtd.amegroups.com/article/view/10.21037/jtd-2025-390/rc
Data Sharing Statement: Available at https://jtd.amegroups.com/article/view/10.21037/jtd-2025-390/dss
Peer Review File: Available at https://jtd.amegroups.com/article/view/10.21037/jtd-2025-390/prf
Funding: This work was supported by
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://jtd.amegroups.com/article/view/10.21037/jtd-2025-390/coif). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). Experiments were performed under a project license (Lunshen No. 230148) granted by the Animal Ethics Committee of The Fourth Affiliated Hospital of Soochow University and in compliance with Soochow university guidelines for the care and use of animals.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Mocumbi A, Humbert M, Saxena A, et al. Pulmonary hypertension. Nat Rev Dis Primers 2024;10:1. [Crossref] [PubMed]
- Li X, Tan J, Wan J, et al. Cell Death in Pulmonary Arterial Hypertension. Int J Med Sci 2024;21:1840-51. [Crossref] [PubMed]
- Novara ME, Di Martino E, Stephens B, et al. Future Perspectives of Pulmonary Arterial Hypertension: A Review of Novel Pipeline Treatments and Indications. Drugs R D 2024;24:13-28. [Crossref] [PubMed]
- Reinders S, Didden EM, Ong R. Survival, morbidity, and quality of life in pulmonary arterial hypertension patients: a systematic review of outcomes reported by population-based observational studies. Respir Res 2024;25:373. [Crossref] [PubMed]
- Simeone B, Maggio E, Schirone L, et al. Chronic thromboembolic pulmonary hypertension: the diagnostic assessment. Front Cardiovasc Med 2024;11:1439402. [Crossref] [PubMed]
- Humbert M, Kovacs G, Hoeper MM, et al. 2022 ESC/ERS Guidelines for the diagnosis and treatment of pulmonary hypertension. Eur Heart J 2022;43:3618-731. [Crossref] [PubMed]
- Chin KM, Channick R, Kim NH, et al. Macitentan and Tadalafil Combination Therapy in Incident and Prevalent Pulmonary Arterial Hypertension: Real-World Evidence from the OPUS/OrPHeUS Studies. Adv Ther 2024;41:4205-27. [Crossref] [PubMed]
- Rodor J, Chen SH, Scanlon JP, et al. Single-cell RNA sequencing profiling of mouse endothelial cells in response to pulmonary arterial hypertension. Cardiovasc Res 2022;118:2519-34. [Crossref] [PubMed]
- Stearman RS, Bui QM, Speyer G, et al. Systems Analysis of the Human Pulmonary Arterial Hypertension Lung Transcriptome. Am J Respir Cell Mol Biol 2019;60:637-49. [Crossref] [PubMed]
- Zheng H, Hua J, Li H, et al. Comprehensive analysis of the expression of N6-methyladenosine RNA methylation regulators in pulmonary artery hypertension. Front Genet 2022;13:974740. [Crossref] [PubMed]
- He Z, Chang T, Chen Y, et al. PARM1 Drives Smooth Muscle Cell Proliferation in Pulmonary Arterial Hypertension via AKT/FOXO3A Axis. Int J Mol Sci 2023;24:6385. [Crossref] [PubMed]
- Dai Z, Thorp EB. New Way to Study Pulmonary Hypertension in HFpEF. Circ Res 2023;133:899-901. [Crossref] [PubMed]
- Gorr MW, Sriram K, Muthusamy A, et al. Transcriptomic analysis of pulmonary artery smooth muscle cells identifies new potential therapeutic targets for idiopathic pulmonary arterial hypertension. Br J Pharmacol 2020;177:3505-18. [Crossref] [PubMed]
- Singh N, Eickhoff C, Garcia-Agundez A, et al. Transcriptional profiles of pulmonary artery endothelial cells in pulmonary hypertension. Sci Rep 2023;13:22534. [Crossref] [PubMed]
- Luo R, Wang T, Lan J, et al. The multifaceted roles of selective autophagy receptors in viral infections. J Virol 2024;98:e0081424. [Crossref] [PubMed]
- Wang L, Zhu Y, Zhang N, et al. The multiple roles of interferon regulatory factor family in health and disease. Signal Transduct Target Ther 2024;9:282. [Crossref] [PubMed]
- Jiang M, Jia K, Wang L, et al. Alterations of DNA damage response pathway: Biomarker and therapeutic strategy for cancer immunotherapy. Acta Pharm Sin B 2021;11:2983-94. [Crossref] [PubMed]
- Sharma BR, Kanneganti TD. Inflammasome signaling in colorectal cancer. Transl Res 2023;252:45-52. [Crossref] [PubMed]
- Zhu KC, Zhang N, Liu BS, et al. Functional Analysis of IRF1 Reveals its Role in the Activation of the Type I IFN Pathway in Golden Pompano, Trachinotus ovatus (Linnaeus 1758). Int J Mol Sci 2020;21:2652. [Crossref] [PubMed]
- Gu T, Lu L, An C, et al. Negative regulation of the RLR-mediated IFN signaling pathway by duck ubiquitin-specific protease 18 (USP18). J Cell Physiol 2019;234:3995-4004. [Crossref] [PubMed]
- Karki R, Sharma BR, Tuladhar S, et al. Synergism of TNF-α and IFN-γ Triggers Inflammatory Cell Death, Tissue Damage, and Mortality in SARS-CoV-2 Infection and Cytokine Shock Syndromes. Cell 2021;184:149-168.e17. [Crossref] [PubMed]
- Chen X, Qi D, Fan S, et al. Interferon regulatory factor 1 (IRF1) inhibits lung endothelial regeneration following inflammation-induced acute lung injury. Clin Sci (Lond) 2023;137:367-83. [Crossref] [PubMed]
- Shen Y, Sun Z, Mao S, et al. IRF-1 contributes to the pathological phenotype of VSMCs during atherogenesis by increasing CCL19 transcription. Aging (Albany NY) 2020;13:933-43. [Crossref] [PubMed]
- Perevalova AM, Gulyaeva LF, Pustylnyak VO. Roles of Interferon Regulatory Factor 1 in Tumor Progression and Regression: Two Sides of a Coin. Int J Mol Sci 2024;25:2153. [Crossref] [PubMed]
- Takahashi K, Sugiyama T, Tokoro S, et al. Inhibitory effect of 10-hydroxydecanoic acid on lipopolysaccharide-induced nitric oxide production via translational downregulation of interferon regulatory factor-1 in RAW264 murine macrophages. Biomed Res 2013;34:205-14. [Crossref] [PubMed]
- Johnson S, Sommer N, Cox-Flaherty K, et al. Pulmonary Hypertension: A Contemporary Review. Am J Respir Crit Care Med 2023;208:528-48. [Crossref] [PubMed]
- Abid S, Marcos E, Parpaleix A, et al. CCR2/CCR5-mediated macrophage-smooth muscle cell crosstalk in pulmonary hypertension. Eur Respir J 2019;54:1802308. [Crossref] [PubMed]
- Wang L, Moonen JR, Cao A, et al. Dysregulated Smooth Muscle Cell BMPR2-ARRB2 Axis Causes Pulmonary Hypertension. Circ Res 2023;132:545-64. [Crossref] [PubMed]
- Liao J, Xie SS, Deng Y, et al. PRDX6-mediated pulmonary artery endothelial cell ferroptosis contributes to monocrotaline-induced pulmonary hypertension. Microvasc Res 2023;146:104471. [Crossref] [PubMed]
- Fan X, Li Q, Wang Y, et al. Non-canonical NF-κB contributes to endothelial pyroptosis and atherogenesis dependent on IRF-1. Transl Res 2023;255:1-13. [Crossref] [PubMed]
- Ferrian S, Cao A, McCaffrey EF, et al. Single-Cell Imaging Maps Inflammatory Cell Subsets to Pulmonary Arterial Hypertension Vasculopathy. Am J Respir Crit Care Med 2024;209:206-18. [Crossref] [PubMed]
(English Language Editor: J. Gray)


