
Relationship between the expression of striated preferentially expressed gene (SPEG) and the development of atrial fibrillation
Highlight box
Key findings
• The striated preferentially expressed gene (SPEG) gene is associated with the incidence of atrial fibrillation, and the common mutation sites of SPEG in the local population are different from those in other parts of the world.
What is known and what is new?
• SPEG has been identified as a critical hub gene in the pathogenesis of atrial fibrillation. SPEG interacts with sarcoplasmic/endoplasmic reticulum Ca2+ ATPase 2a (SERCA2a) to promote the movement of calcium ions.
• In this study, SPEG expression differed between the atrial fibrillation and sinus rhythm groups. Receiver operating characteristic curve analysis indicated that SPEG expression had a high accuracy (area under the curve >0.90) in predicting atrial fibrillation. Atrial fibrillation hub genes are involved in positive regulation of transcription by RNA polymerase II. The presence of a rare G/A mutation in SPEG at the RS576016632 locus patients with atrial fibrillation from Guangxi differs from the global data of the Human Genome Project.
What is the implication, and what should change now?
• The abnormal expression of SPEG was positively correlated with the incidence of atrial fibrillation.
• The expression of SPEG varies significantly across individuals as does the pathogenesis of atrial fibrillation associated with it.
• Carefully screening of other loci for the SPEG gene among the Guangxi population should be conducted to determine the gene frequency in this region.
Introduction
Atrial fibrillation (AF) is a commonly observed type of rapid atrial arrhythmia in clinical practice, which often results in severe complications such as heart failure and cardiogenic embolism (1). AF can cause changes in cardiac function, leading to hemodynamic disturbances, and the left ventricular ejection fraction in patients with AF is approximately 25% lower than that in healthy individuals. Moreover, patients with AF exhibit reduced atrial filling or increased pressure in both atria, resulting in a higher incidence of arrhythmogenic cardiomyopathy (2-4). AF has also been linked to extracardiac diseases (1). With the advancements made in basic and clinical research on AF over the past few decades, various risk factors have been identified, including smoking, hypertension, obesity, diet, age, and sex. Moreover, genetic risk factors have garnered increased attention (2). However, AF is a complex polygenic condition, and its pathogenesis remains unclear (5).
AF can be categorized into valvular AF (VAF) and nonvalvular AF (NVAF). The pathogenesis of AF involves both the initiation and maintenance of pathogenic mechanisms. Genetic mutations influence the occurrence and progression of AF to a certain extent (6,7). Research has identified the crucial role of ion channels in AF pathogenesis (8). Therefore, genetic mutations that affect ion channel structures have become a focus of research (9). Recently, various genes and factors related to calcium ion channels have been identified. The early release of calcium stored in the sarcoplasmic reticulum may be related to the occurrence and maintenance of AF. In patients with chronic AF, fibrous scars formed by greater fibrosis can also be seen (10). Striated preferentially expressed gene (SPEG), a critical regulator of calcium ions, contributes to the occurrence and maintenance of AF by influencing the duration of action potentials (11,12).
AF is the result of multigene synergism. First, a recent study has shown that SPEG is involved in the formation of a tripartite structure composed of sarcoplasmic reticulum, transverse striae, and other components, which plays a key role in the transport of calcium ions in cardiomyocytes, actively moving calcium ions into the sarcoplasmic reticulum for storage (4). Second, SPEG is also highly expressed in the mitochondria of muscle cells, with the SPEG alpha subunit being specifically expressed in cardiomyocytes. Abnormalities, namely single-nucleotide polymorphisms (SNPs) and single-nucleotide variations (SNVs), can also cause mitochondrial ridge structure abnormalities and a decrease in the mitochondrial ATP production rate (13).
In this study, we analyzed the GSE41177 and GSE128188 datasets from the Gene Expression Omnibus (GEO) database and known human pathogenic genes related to AF obtained from the GeneCards database (http://www.genecards.org/) in order to confirm the involvement of SPEG in AF pathogenesis. Subsequently, using bioinformatic tools such as RStudio, Cytoscape software, and the Microbioinformatics website (http://www.bioinformatics.com.cn/), we aimed to screen, analyze, and summarize the expression of SPEG in the myocardial tissue of individuals with AF and those with sinus rhythm (SR). Additionally, the relationship between SPEG and AF was clarified through receiver operating characteristic (ROC) curve analysis. Finally, by screening mutation sites in the Human Genome Project we identified exons harboring AF-associated SNPs. Samples were collected from 129 patients with NVAF, 60 patients with VAF, and 100 individuals with SR in the Guangxi region. Exon sequencing was then conducted to investigate the mutation status of the RS576016632 locus of SPEG in the NVAF, VAF, and SR groups. We present this article in accordance with the STREGA reporting checklist (available at https://jtd.amegroups.com/article/view/10.21037/jtd-2025-456/rc).
Methods
Study population and data acquisition
Using the search term “atrial fibrillation”, we retrieved two datasets, GSE41177 and GSE128188, from the GEO database (www.ncbi.nlm.nih.gov/geo). The GSE41177 dataset consisted of gene expression profile data from 32 atrial tissue samples in the AF group and six atrial tissue samples in the SR group. Meanwhile, the GSE128188 dataset comprised gene expression profile data from 10 atrial tissue samples in the AF group and four atrial tissue samples in the SR group. Detailed information about these datasets is presented in Table 1.
Table 1
| SNP | Location | Mutation type | Mutation site | Amino acid site |
|---|---|---|---|---|
| RS576016632 | Coding region | Missense mutation | G/A | NP:005867.3:p.Arg5Leu |
To determine the specific location and obtain details of the SPEG mutations at the RS576016632 site, we analyzed available information on the UniProt and NCBI websites. SNP, single-nucleotide polymorphism; SPEG, striated preferentially expressed gene; NCBI, National Center for Biotechnology Information.
Differential gene expression analysis
Gene expression matrices of the GSE41177 and GSE128188 datasets were downloaded and subjected to differential gene expression analysis using the “GEOquery” and “Limma” packages in the R version 4.2.2 software (RStudio). In the GSE41177 dataset, differentially expressed genes (DEGs) were identified using a threshold of P value <0.05 and |log2 fold change| >0.50. In the GSE128188 dataset, DEGs were identified using a threshold of P<0.05 and |log2 fold change| >0.40. Subsequently, differential gene expression data were exported and visualized as volcano plots and heatmaps using the Microbioinformatics online website, with the target genes being highlighted.
Identification of potential AF-related target genes
Known AF-related human genes were identified and downloaded from the GeneCards database using the term “atrial fibrillation”. Genes with a relevance score of >2 in each dataset were selected as the AF-related genes. Subsequently, Venn diagrams of the DEGs from the GSE41177 and GSE128188 datasets intersecting with the genes obtained from the GeneCards database were plotted. The overlapping genes were considered potential AF target genes and further analyzed.
Protein-protein interaction (PPI) analysis of the potential target genes and identification of hub genes
To evaluate protein interactions among the potential target genes, we employed the Search Tool for the Retrieval of Interacting Genes (STRING; version 12.0; https://cn.string-db.org/). First, potential target genes were imported into STRING to construct a PPI network. The confidence threshold (medium confidence) was set to 0.15. Subsequently, the corresponding results were imported into the Cytoscape software (version 3.10.0) for hub gene screening and visualization. Through the CytoNCA plugin in Cytoscape, betweenness centrality (BC), closeness centrality, and degree centrality were selected as the main criteria, with BC being the primary selection standard. The top 50 genes under these three criteria were identified as hub genes.
Functional enrichment analysis of hub genes
Gene Ontology (GO) is a framework for the functional annotation of genes, categorizing them into biological process (BP), cellular components (CC), and molecular function (MF). Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis is the most common approach employed for pathway annotation. In this study, hub genes were imported into the Database for Annotation, Visualization and Integrated Discovery (DAVID) online database (version 6.8; https://david.ncifcrf.gov/summary.jsp) for analysis. The top 10 entries with P<0.05 were selected for visualization via the Microbioinformatics website.
Analysis of SPEG expression
The expression level of SPEG in both the AF and SR groups was analyzed using the validation dataset GSE41177, and the results were visualized using the “ggpubr” package in RStudio. In addition, the Human Protein Atlas online database (http://www.proteinatlas.org/) was used to analyze the expression level of SPEG in various tissues. This database provides information on the expression of protein-coding genes in all major human tissues and organs.
Analysis of the diagnostic value of SPEG
The GSE41177 dataset was used to validate expression levels of SPEG. After the expression data for each sample were downloaded, the software (IBM Corp., Armonk, NY, USA) was employed to perform ROC curve analysis, which enabled determination of the specificity, positive predictive value, sensitivity, likelihood ratio, and negative predictive value of SPEG at all possible thresholds. Subsequently, the accuracy of SPEG in predicting AF susceptibility was evaluated based on the results of ROC curve analysis.
Selection and sequencing of target gene loci
Next, the SPEG gene loci were searched in the SNP database on the PubMed website (https://www.ncbi.nlm.nih.gov/snp/). The criteria for SNP site selection were as follows: (I) sites located in functional regions of the gene, such as the promoter, coding, and exon regions; (II) sites with a minor allele frequency of ≥2%; and (III) sites reported to be associated with AF susceptibility in the literature. Based on these criteria, the RS576016632 locus was identified.
We selected 129 patients with NVAF and 60 patients with VAF who had been admitted to the Department of Cardiovascular Medicine and Cardiothoracic Surgery at the Affiliated Hospital of Youjiang Medical University for Nationalities and Wuzhou People’s Hospital from November 2021 to February 2024 to form the case group. In addition, 100 individuals diagnosed with SR at the Physical Examination Center of both hospitals in the same period were selected as the control group. The general clinical data of patients age, creatine kinase MB isoenzyme (CK-MB), partial thromboplastin time (PTT), international normalized ratio (INR), activated partial thromboplastin time (APTT), creatinine (CREA), left atrial end-systolic transverse diameter (LAsD), right atrial end-systolic transverse diameter (RAsD), ejection fraction (EF), sex, cardiac troponin I (cTnI), myoglobin (MYO), and platelet (PLT) were collected for statistical analysis. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). This study received approval from the Ethics Committees of the Affiliated Hospital of Youjiang Medical University for Nationalities (No. YYFY-LL-2023-173) and Wuzhou People’s Hospital (No. 2022093). All participants and their families provided written informed consent.
Experimental and sequencing methods
Morning fasting venous blood samples (4 mL) were collected from the study participants into anticoagulant tubes containing EDTA K2. After labeling, all samples were stored at −80 ℃ until analysis. Genomic DNA was then extracted using a magnetic bacterium genomic DNA kit (Xi’an Biolab Biotechnology Co., Ltd., Xi’an, China). After extraction, the quantity and quality of DNA were determined using agarose electrophoresis. The samples were then stored at −80 ℃ until sequencing. Subsequently, the extracted DNA was sent to the Wuhan Branch Office of Beijing Genomics Institute for locus DNA sequencing of SPEG via the Sanger sequencing method to identify mutations in the corresponding samples. The primer sequences used for the amplification and sequencing are shown in Table S1. We randomly selected 10 samples from the AF group and the control group and sent them to Beijing Genomics Institution (Wuhan, China) for quantitative polymerase chain reaction (qPCR) detection. We also analyzed SPEG gene expression in the samples of the two groups.
Statistical analysis
All statistical analyses were performed using the SPSS 19.0 software package (IBM Corp.). The clinical data were all in line with the normal distribution detected by SPSS 19.0. Measurement data were further tested by the independent samples t-test for specificity, and count data were tested via Chi-squared tests. The results of gene expression analysis of the GSE41177 and GSE128188 datasets were consistent with positive distribution and were further analyzed via the t-test. Cluster analysis of BP, MF, and CC terms was performed, and the hypergeometric P value was calculated. The chi-squared test was used for determining Hardy-Weinberg equilibrium of count data and genotype. Results with a P<0.05 were considered statistically significant.
Results
Volcano plot and heatmaps of DEGs
In this study, we identified a total of 19,540 DEGs between the AF and SR control groups in the GSE41177 dataset. Of these, 19,531 genes were upregulated, and 9 genes were downregulated in the AF group. A volcano plot of differential gene expression is shown in Figure 1A, and a heatmap of the top 50 DEGs is displayed in Figure 1B. Meanwhile, there were 2,258 DEGs in the GSE128188 dataset between the AF and SR groups, of which 918 were upregulated and 1,340 were downregulated. A volcano plot of the differential gene expression profile is shown in Figure 1C, and a heatmap of the top 50 DEGs is shown in Figure 1D.
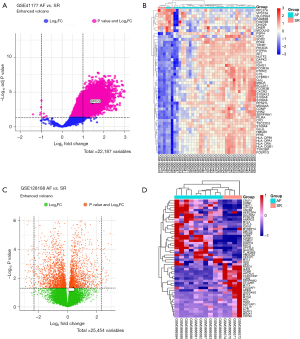
Screening of potential target genes
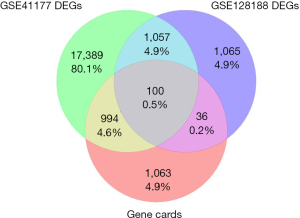
Screening of the GeneCards database revealed 5,076 genes associated with AF, of which 2,193 genes had a relevance score of >2. Notably, a total of 100 DEGs were shared across the GSE41177, GSE128188, and GeneCards datasets. These were selected for further analysis. Among the 100 overlapping genes, SPEG was identified as a potential target (Figure 2).
PPI network analysis and hub gene screening
The 100 intersecting genes were imported into the STRING database for PPI network analysis. After the removal of isolated nodes, the constructed PPI network included 95 nodes (proteins) and 1,165 edges (PPIs), with an average node degree of 24.5 and a PPI enrichment P value of <0.001. Subsequently, the corresponding .TSV file was exported and imported into Cytoscape software. Topological calculations were performed with the CytoNCA plugin, and the top 60 genes ranked by BC were considered hub genes. The hub genes included MYH7, JUN, VWF, MYL3, NPPA, CACNB2, KCNE4, SYNE2, SPEG, and KCNAB2 (Figure 3).
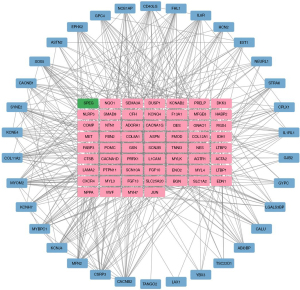
GO and KEGG enrichment analyses
KEGG pathway enrichment results are presented as a bubble chart in Figure 4A. Hub genes were primarily enriched in the cAMP signaling, cardiac muscle contraction, cyclic guanosine monophosphate-protein kinase G (cGMP-PKG) signaling, calcium signaling, and apelin signaling pathways. The GO enrichment analysis results, including BP, CC, and MF terms, are displayed in bar charts. As shown in Figure 4B, the hub genes were predominantly associated with the following BPs: cell adhesion, regulation of blood pressure, calcium-mediated signaling, cell-cell signaling, positive regulation of heart rate, and positive regulation of ATP-dependent activity.
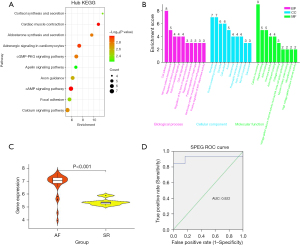
With regard to CC terms, hub genes were primarily enriched in the synapse, protein-containing complex, dendrite, cell surface, axon, sarcolemma, presynaptic membrane, actin cytoskeleton, myosin II complex, and sarcomere.
Regarding MF terms, hub genes were primarily enriched in calcium ion binding, heparin binding, actin binding, hormone activity, actin filament binding, collagen binding, voltage-gated calcium channel activity involved SA node cell action potential, myosin II heavy chain binding, microfibril binding, and high voltage-gated calcium channel activity.
SPEG expression in various tissues
The GSE41177 dataset, comprising 32 AF samples and 6 SR samples, was used as the validation dataset. Notably, the AF group exhibited significantly higher SPEG expression than did the SR group (P<0.001), as shown in Figure 4C.
ROC curve analysis of SPEG expression
Using the GSE41177 dataset, ROC curve analysis was performed to evaluate the accuracy of SPEG in predicting AF susceptibility. SPEG expression was significantly higher in the AF group than in the SR group in the validation dataset (P<0.001). Additionally, the area under the ROC curve (AUC) for SPEG expression was 0.92, indicative of high accuracy in predicting AF risk (Figure 4D).
Patient data
By comparing the clinical data of patients with the RS576016632 mutation in the AF group to that of patients without the mutation, we found statistically significant differences in age, PTT, INR, APTT, LAsD, RAsD, EF (P<0.001), CREA (P=0.03) and CK-MB (P=0.006). There was no significant difference in gender (P=0.35), cTnI (P=0.24), MYO (P=0.09) and PLT (P=0.07) (Table 2).
Table 2
| Variables | t/χ2† | P value |
|---|---|---|
| Age | 7.927 | <0.001 |
| Sex (male/female) | 0.865 | 0.35 |
| cTnI (µg/mL) | −1.192 | 0.24 |
| MYO (µg/mL) | −1.706 | 0.09 |
| CK-MB (U/L) | −2.754 | 0.006 |
| PTT (s) | 7.092 | <0.001 |
| INR | 7.560 | <0.001 |
| APTT (s) | 4.548 | <0.001 |
| CREA (µmol/L) | 2.201 | 0.03 |
| PLT (×109) | −1.795 | 0.07 |
| Echocardiography | ||
| LAsD (mm) | 15.285 | <0.001 |
| RAsD (mm) | 9.388 | <0.001 |
| EF (%) | −4.287 | <0.001 |
†, age and sex are χ2 values, and the others are t values. cTnI, cardiac troponin I; MYO, myoglobin; CK-MB, creatine kinase MB isoenzyme; PTT, partial thromboplastin time; INR, international normalized ratio; APTT, activated partial thromboplastin time; CREA, creatinine; PLT, platelet; LAsD, left atrial end-systolic transverse diameter; RAsD, right atrial end-systolic transverse diameter; EF, ejection fraction.
The RS576016632 locus is situated within the coding region of SPEG (Table 1). This mutation is a G>A substitution, resulting in a change of arginine with leucine in the protein product, causing a substantial alteration in protein structure. To determine the specific location and obtain details on the rs576016632 variant, the UniProt (https://www.uniprot.org) and National Center for Biotechnology Information (NCBI) (https://www.ncbi.nlm.nih.gov/gap/phegeni) databases were used (Table 3). The genotype distribution of rs576016632 was examined in 129 patients with NVAF, 60 patients with VAF, and 100 control individuals from the Guangxi region. The rs576016632 SPEG locus was sequenced in 189 patients with AF and 100 controls with SR without the mutation. The results of statistical analysis were significantly different compared to those of the Human Genome Project (P=0.049).
Table 3
| Region | No. of cases | Allele, n (%) | |||
|---|---|---|---|---|---|
| G | A | T | C | ||
| Guangxi | 189 | 378 (100.00) | 0 | 0 | 0 |
| World | 52,087 | 104,157 (99.98) | 1 (<0.001) | 15 (<0.001) | 1 (<0.001) |
| South Africa | 661 | 1,322 (100.00) | 0 | 0 | 0 |
| United States | 346 | 691 (99.99) | 1 (0.01) | 0 | 0 |
| Spain | 504 | 1,008 (100.00) | 0 | 0 | 0 |
| Europe | 7,698 | 15,395 (100.00) | 0 | 0 | 1 (<0.001) |
| Latin America | 10,700 | 21,385 (99.99) | 0 | 15 (<0.001) | 0 |
| P value | 0.049 | ||||
G, guanine; A, adenine; T, thymine; C, cytosine.
Genotypes of the rs576016632 locus in 189 samples sequenced in the study region’s population were compared with single-nucleotide distributions in the global population and populations from South Africa, the United States, Spain, Europe, and Latin America.
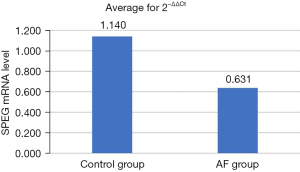
In order to assess SPEG gene expression in the control group and the AF group, we selected 10 samples from each group for qPCR. Compared with the control group, SPEG messenger RNA (mRNA) expression was significantly decreased in the AF group (Figure 5).
Discussion
In this study, we found that SPEG expression was upregulated in AF through bioinformatic analysis. Additionally, we demonstrated that SPEG could accurately predict the risk of AF. Moreover, SPEG was found to correlate with ion channel function. AF is considered to be a complex condition marked by impairments in the electrophysiological activity of the heart and is caused by various factors. Pathological changes, such as abnormal ion channel function and myocardial fibrosis, can promote and maintain the onset of AF, with common calcium, potassium, and sodium pumps commonly being affected (14,15). The current treatment strategies focus on controlling the ventricular rate or restoring AF to SR. However, owing to the high recurrence rate of AF and the adverse effects of these treatments, investigations into new therapeutic approaches are underway. There has been a recent shift in research toward targeted therapies, including pathway- and substrate-specific agents (upstream therapies) (16,17).
Abnormal calcium concentration in the sarcoplasmic reticulum and muscular duct system is closely related to the onset of AF due to abnormal coding of calcium ion channels (18). Mutations in genes encoding ion channels and abnormalities in related accessory proteins and factors contribute to the emergence and persistence of AF (19,20). Determining the molecular mechanisms involved in the pathogenesis of AF has facilitated the development of effective treatments. In recent years, multiple risk factors for AF have been identified, including a growing number of genetic mutations and polymorphisms (21).
In this study, 72 AF-related genes were identified from the GEO and the GeneCards databases through bioinformatic analysis, and 50 core genes were selected based on node centrality via a protein interaction website. GO and KEGG enrichment analyses revealed that calcium channels play a key role in AF pathogenesis, which is consistent with the findings of a previous study (6). We subsequently examined the relationship between the key SPEG gene and AF development in relation to calcium homeostasis. The RS576016632 G/A mutation locus was screened using the NCBI database to assess the risk of AF in patients living in Guangxi.
In recent years, various bioinformatic methods have emerged, but only a few have been widely adopted in China, which may be attributed to the limitations in the capabilities of existing methods to appropriately account for significant ethnic diversity, disease specificity, and differences in living habits across patient populations. As common bioinformatic tools, such as Regmex, require further validation in this population, we did not use Regmex or other methods for identifying important motifs in sequence lists, nor did we conduct motif analysis or traditional k-mer screening. We screened the local population by sequencing exon loci with high mutation frequency in the Human Genome Project.
Genome-wide association studies have identified several AF susceptibility-related mutation loci, specifically in genes associated with calcium channels, and genes upstream and downstream of SPEG have been examined to determine whether there is a synergistic effect. SYNPO2L8 and TTN are involved in atrial myocardial remodeling, which leads to AF in cardiomyopathy (22). Mutations in SPEG (linked to myocyte diseases) that are associated with dilated cardiomyopathy and cardiomyopathy-induced AF have been identified (22,23). Moreover, TTNtv has been found to be significantly enriched in familial and early-onset AF. In one study, the genetic risk in the population of British residents of European descent was assessed based on predicted loss of function variants and polygenic risk (24). However, there was no obvious familial clustering of AF in the population within the study area. Our study did not involve family-based research or multimethod susceptibility assessments. Instead, we focused on the role SPEG plays in the electromagnetic homeostasis related to AF pathogenesis and examined the rs576016632 G/A mutation in patients with AF.
The SPEG alpha subunit is overexpressed in cardiomyocytes, promoting calcium channel maturation via ridge development and phosphorylation. Abnormal expression of the SPEG gene causes mitochondrial dysfunction, characterized by significantly reduced ATP production and a marked decrease in the concentration of calcium ions in the sarcoplasmic reticulum. An unstable calcium concentration is closely related to electrical activity in AF. Factors upstream and downstream of SPEG, such as RyR2, also play a critical role in the development of AF, contributing to a decline in sarcoplasmic reticulum function through phosphoric acid deficiency (10). Conversely, mutations in this gene or reduction in its expression can lead to an abnormal increase in calcium ion release, causing delayed afterdepolarizations (DADs) with increased amplitude, which upon reaching the threshold, can trigger AF. The generation of early afterdepolarizations (EADs) and DADs results in shortened action potentials. In addition, phosphorylation of the SERCA2a receptor at site T484 increases calcium uptake. Notably, abnormal enhancement of SERCA2a receptor activity can accelerate Ca2+ cycling, leading to faster production of Ca2+ transients and increased activation of sodium-calcium exchangers. This results in membrane depolarization and a reduced DAD threshold, thereby increasing the likelihood of AF development. Stabilizing the junctional membrane complex and ensuring the structural integrity of the JPH-2 protein is essential for effective electrical conduction. Structural disruption can lead to electrical disturbances, promoting the occurrence and maintenance of AF (6,11,17). Meanwhile, animal experiments have revealed that greater cardiac interstitial fibrosis with abnormal SPEG expression. The relationship between fibrosis, membrane potential, and action potential requires further study (13). The rs57601662 SPEG variant is a missense mutation, located in the coding region at Chr2:19434991. This mutation, which affects the protein backbone, is a substitution of G with A, resulting in a change of arginine to leucine (5). Ultimately, this alteration changes SPEG protein structure, thereby affecting cellular functions and potentially causing AF. In this study, the G/A mutation at this locus found not to be a significant factor in AF development. Bioinformatic analysis of screen data from Taiwan and London revealed a close relationship between SPEG expression and AF development. The lack of G/A mutations in the Guangxi region may be attributed to geographic factors. This region is located in southwestern China, bordering several Southeast Asian countries. It is characterized by a subtropical monsoon climate and comprises various ethnic groups, including Han, Zhuang, Bouyei, Miao, Yao, and Shui. The long-term economic underdevelopment and low population mobility within this region make its genetic pool relatively stable and representative. We sequenced the loci of the target gene SPEG. Target sequences are often regulated by cis-acting and trans-acting elements in noncoding sequences, which can activate, inhibit, or silence gene expression (25). The results in our study may be attributed to the suppression of the promoters or transcription factors upstream of the mutation sites, resulting in downregulated expression.
As one of the common arrhythmia diseases, there are many biomarkers for atrial fibrillation. In this study, the author tried to use SPEG gene as one of the biomarkers for atrial fibrillation. However, the specific role of SPEG gene still needs further study. In this paper, we analyzed the common biomarkers of atrial fibrillation. By comparing a current study, more than dozens of clear markers of atrial fibrillation have been identified (e.g., leptin, adiponectin, ceramide, ferritin, fibrinogen, interleukin-18, thromboreactive protein-1, acylcarnitine, plasminogen activator inhibitor-1, triglycerides, high-density lipoprotein cholestasis, intestinal microbiome, hyperuricemia, and homocysteine) (26). However, at present, there is no direct interaction between the above markers and the SPEG gene studied in this paper. In order to clarify the role of SPEG as a biomarker for atrial fibrillation, the author of this paper will further study the mechanism of SPEG gene action.
Finally, data mining and analysis revealed a correlation between SPEG expression and AF development. The GEO data indicated that SPEG expression is upregulated in patients with AF as compared with that in individuals with SR. In addition, statistical analysis revealed SPEG expression in the heart to be a highly accurate predictor of AF risk, indicating a close association between SPEG expression and AF pathogenesis. Subsequently, the rs576016632 locus was screened to further clarify the association of SPEG with AF. Although our results indicated a negative association between the rs576016632 variant and AF, this finding does not necessarily rule out a correlation between SPEG expression and AF pathogenesis as our study involved a small sample, suboptimal sample selection, and incomplete evaluation of mutation sites. Overall, the findings of this study may not be fully representative, and future studies with larger samples are required to investigate other notable loci to further validate the relationship between SPEG and NVAF.
This study involved other limitations. Specifically, our investigation relied solely on bioinformatic approaches, and we did not conduct corresponding validation experiments in vitro or in vivo. Therefore, further research should focus on validation experiments to confirm the conclusions of this study.
Conclusions
SPEG is critically involved in maintaining the stability of electrical conduction mechanisms and calcium homeostasis. The failure to effectively promote the maturation of calcium ion channels leads to abnormal calcium ion concentration in the sarcoplasmic reticulum, which is involved in the formation and persistence of AF (11,27). Furthermore, our bioinformatic analysis demonstrated that the hub genes associated with AF may play key roles in the development of AF through various pathways and processes, including cardiac conduction regulation of heart rate, calcium ion transmembrane transport, voltage-gated calcium channel complex formation, MAPK signaling, cAMP signaling, and calcium signaling. These findings align with those of previous studies (15,28-30). Moreover, rs576016632 variant was not detected in this study, and SPEG gene expression in the AF group was lower than that in SR group, but the difference between the two was not statistically significant. Our results might have been biased by the small sample size and population specifics in the geographic region, and thus further research is needed.
Acknowledgments
We would like to thank Editage (www.editage.cn) for the English language editing.
Footnote
Reporting Checklist: The authors have completed the STREGA reporting checklist. Available at https://jtd.amegroups.com/article/view/10.21037/jtd-2025-456/rc
Data Sharing Statement: Available at https://jtd.amegroups.com/article/view/10.21037/jtd-2025-456/dss
Peer Review File: Available at https://jtd.amegroups.com/article/view/10.21037/jtd-2025-456/prf
Funding: This work was supported by
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://jtd.amegroups.com/article/view/10.21037/jtd-2025-456/coif). B.D. reports funding, the provision of study materials, medical writing, and support for article processing charges from the Project of Improving the Basic Scientific Research Ability of Young College Teachers in Guangxi (No. 2023KY2019), and the 2022 Wuzhou City Health Commission Scientific Research Project (No. WZWS-F2022012). Both funds supported genetic sequencing. The other authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). This study received approval from the Ethics Committees of the Affiliated Hospital of Youjiang Medical University for Nationalities (No. YYFY-LL-2023-173) and Wuzhou People’s Hospital (No. 2022093). All participants and their families provided written informed consent.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Elsheikh S, Hill A, Irving G, et al. Atrial fibrillation and stroke: State-of-the-art and future directions. Curr Probl Cardiol 2024;49:102181. [Crossref] [PubMed]
- Gondal MUR, Mehmood RS, Khan RP, et al. Atrial myopathy. Curr Probl Cardiol 2024;49:102381. [Crossref] [PubMed]
- Hu D, Barajas-Martinez H, Zhang ZH, et al. Advances in basic and translational research in atrial fibrillation. Philos Trans R Soc Lond B Biol Sci 2023;378:20220174. [Crossref] [PubMed]
- Elliott AD, Middeldorp ME, Van Gelder IC, et al. Epidemiology and modifiable risk factors for atrial fibrillation. Nat Rev Cardiol 2023;20:404-17. [Crossref] [PubMed]
- Khawajakhail R, Khan RU, Gondal MUR, et al. Advancements in gene therapy approaches for atrial fibrillation: Targeted delivery, mechanistic insights and future prospects. Curr Probl Cardiol 2024;49:102431. [Crossref] [PubMed]
- Rennison JH, Van Wagoner DR. Dysregulated Ca2+ cycling in atrial fibrillation. Eur Heart J 2023;44:2495-7. [Crossref] [PubMed]
- Okamura S, Onohara Y, Ochi H, et al. Minor allele of GJA1 gene polymorphism is associated with higher heart rate during atrial fibrillation. Sci Rep 2021;11:2549. [Crossref] [PubMed]
- Colman MA, Varela M, MacLeod RS, et al. Interactions between calcium-induced arrhythmia triggers and the electrophysiological-anatomical substrate underlying the induction of atrial fibrillation. J Physiol 2024;602:835-53. [Crossref] [PubMed]
- Jameson HS, Hanley A, Hill MC, et al. Loss of the Atrial Fibrillation-Related Gene, Zfhx3, Results in Atrial Dilation and Arrhythmias. Circ Res 2023;133:313-29. [Crossref] [PubMed]
- Li G, Huang H, Wu Y, et al. Striated preferentially expressed gene deficiency leads to mitochondrial dysfunction in developing cardiomyocytes. Basic Res Cardiol 2024;119:151-68. [Crossref] [PubMed]
- Lee CS, Jung SY, Yee RSZ, et al. Speg interactions that regulate the stability of excitation-contraction coupling protein complexes in triads and dyads. Commun Biol 2023;6:942. [Crossref] [PubMed]
- Heijman J, Zhou X, Morotti S, et al. Enhanced Ca(2+)-Dependent SK-Channel Gating and Membrane Trafficking in Human Atrial Fibrillation. Circ Res 2023;132:e116-33. [Crossref] [PubMed]
- Li Q, Lin J, Luo S, et al. Integrated multi-omics approach reveals the role of striated muscle preferentially expressed protein kinase in skeletal muscle including its relationship with myospryn complex. J Cachexia Sarcopenia Muscle 2024;15:1003-15. [Crossref] [PubMed]
- Le Quilliec E, LeBlanc CA, Neuilly O, et al. Atrial cardiomyocytes contribute to the inflammatory status associated with atrial fibrillation in right heart disease. Europace 2024;26:euae082. [Crossref] [PubMed]
- Voigt N, Seibertz F, Fakuade FE. Vicious Twins: RyR2 Dysfunction and Structural Remodeling. Circ Res 2023;133:193-5. [Crossref] [PubMed]
- Wang N, Yu Y, Sun Y, et al. Acquired risk factors and incident atrial fibrillation according to age and genetic predisposition. Eur Heart J 2023;44:4982-93. [Crossref] [PubMed]
- Gillis AM, Dobrev D. Targeting the RyR2 to Prevent Atrial Fibrillation. Circ Arrhythm Electrophysiol 2022;15:e011514. [Crossref] [PubMed]
- Campbell HM, Quick AP, Abu-Taha I, et al. Loss of SPEG Inhibitory Phosphorylation of Ryanodine Receptor Type-2 Promotes Atrial Fibrillation. Circulation 2020;142:1159-72. [Crossref] [PubMed]
- Benitah JP, Perrier R, Mercadier JJ, et al. RyR2 and Calcium Release in Heart Failure. Front Physiol 2021;12:734210. [Crossref] [PubMed]
- Munro ML, van Hout I, Aitken-Buck HM, et al. Human Atrial Fibrillation Is Not Associated With Remodeling of Ryanodine Receptor Clusters. Front Cell Dev Biol 2021;9:633704. [Crossref] [PubMed]
- Babini H, Jiménez-Sábado V, Stogova E, et al. hiPSC-derived cardiomyocytes as a model to study the role of small-conductance Ca(2+)-activated K(+) (SK) ion channel variants associated with atrial fibrillation. Front Cell Dev Biol 2024;12:1298007. [Crossref] [PubMed]
- Vad OB, Monfort LM, Paludan-Müller C, et al. Rare and Common Genetic Variation Underlying Atrial Fibrillation Risk. JAMA Cardiol 2024;9:732-40. [Crossref] [PubMed]
- Ahlberg G, Refsgaard L, Lundegaard PR, et al. Rare truncating variants in the sarcomeric protein titin associate with familial and early-onset atrial fibrillation. Nat Commun 2018;9:4316. [Crossref] [PubMed]
- Nielsen MM, Tataru P, Madsen T, et al. Regmex: a statistical tool for exploring motifs in ranked sequence lists from genomics experiments. Algorithms Mol Biol 2018;13:17. [Crossref] [PubMed]
- Yang JH, Hansen AS. Enhancer selectivity in space and time: from enhancer-promoter interactions to promoter activation. Nat Rev Mol Cell Biol 2024;25:574-91. [Crossref] [PubMed]
- Rafaqat S, Radoman Vujačić I, Gluscevic S, et al. Biomarkers of diabetes: role in the pathogenesis of atrial fibrillation. Eur Rev Med Pharmacol Sci 2024;28:1524-40. [Crossref] [PubMed]
- Campbell H, Aguilar-Sanchez Y, Quick AP, et al. SPEG: a key regulator of cardiac calcium homeostasis. Cardiovasc Res 2021;117:2175-85. [Crossref] [PubMed]
- Tarifa C, Vallmitjana A, Jiménez-Sábado V, et al. Spatial Distribution of Calcium Sparks Determines Their Ability to Induce Afterdepolarizations in Human Atrial Myocytes. JACC Basic Transl Sci 2023;8:1-15. [Crossref] [PubMed]
- Duan S, Du J. Sinus node dysfunction and atrial fibrillation-Relationships, clinical phenotypes, new mechanisms, and treatment approaches. Ageing Res Rev 2023;86:101890. [Crossref] [PubMed]
- Pabel S, Knierim M, Stehle T, et al. Effects of Atrial Fibrillation on the Human Ventricle. Circ Res 2022;130:994-1010. [Crossref] [PubMed]
(English Language Editor: J. Gray)


